Debasis Mitra
Florida Tech School of Computing, Professor
Currently a Professor at the Florida Institute of Technology Debasis Mitra is a Physicsist and a Computer Scientist.
A central theme of Mitra's present research is to squeeze knowledge nuggets from noisy spatio-temporal data, especially in Bio-medical images. One of these efforts is dedicated to improving reconstruction of Nuclear Medical images of heart and brain. A long term vision is to understand the molecular biology of deseases for improving human health. In the past Mitra has made contributions in artificial intelligence studying the qualitative reasoning problems with spatial and temporal constraints.
Pseudo Heart.pptx(just in case: PseudoHeart.pdf)
Courses
I learn, along with my students.
- Fall 2024: Artificial Intelligence
- Spring 2018: Topological Data Analyses
- Spring 2017 (Fall 2022): Scientific Computing
- Past Course: Formal Languages and Automata Theory
- Past Course: Analysis of Algorithms
- Past Course: Medical Imaging
- Past Course: Computational Molecular Biology
- Past Course: Constraint Reasoning
- Future Course: Topological Data Analyses
Approved Humanities Electives for Spring2015
Approved Science Electives for Spring2015
Restricted Business Electives 2015
(Disclaimer: I received them in 2015. The list is not necessearily accurate/"official"/mandatory, it is rather a suggestive one)
On a: learning contract
Stanford class-central
Stanford coursera
MIT mitx
MIT udacity
Primary question my lab asks is given sequence of images acquired at different points in time and perspective, or spatio-temporal data/image, where we have some preliminary model, how much information can we extract from it? Apart from oerall statistical model of noise, we often have extra information about what we are looking for. Can we embed such knowldge in the processing pipeline to go beyond the information content in the data? In essence, this is combined output of data and qualitative knowledge on the target objective.
Message for prospective students
A talk on Medical imaging
Graduate Students
- Valerie Kobzarenko, Ph.D. Candidate
Alumni (Dissertation/Thesis/Capstone)
- Haoran Chang, Ph.D., Post-doc Univ California, Davis
- Michael Kolar, M.S., Medtronic, Boston
- Gengbo Liu, Ph.D., , Genentech Corp., San Francisco
- Haoran Chang, M.S., currently doing Ph.D. with me at FIT
- Hui Pan, Ph.D., TenCent, China, previously at Microsoft Corp.
- Mahmoud Abdalah, Ph.D., Research Associate, Moffitt Cancer Research Center, U of South Florida, Tampa
- Stephen Johnson, Ph.D., Dean, East Florida State College, Palm Bay, Florida
- Chen Shi, M.S., Stanley Black & Decker, Atlanta, GA
- Bo Li, M.S.
- Kimberley Day, B.S., Google Corp.
- Daniel Eiland, M.S., previously at Harris Corp.
- Antall Fernandes, M.S., Visible Measures Corp, Boston
- Richard Hoch, M.S., previously at General Dynamics
- Abhishek, Bannerjee, B.Tech.-IIT, Google, India
- Florent Launay, M.S., Microsoft Corp.
- Sung Park, M.S., Microsoft Corp.
- Keith Ledig, M.S., Harris Corp.
- Gandhali Samanth, M.S., Microsoft Corp.
- Michael Smith, M.S., (somewhere in New York!)
Alan Bundy's Page on how-to-do research
Collaborators
- Grant Gullberg & Group, Department of Radiotracer Development and. Imaging Technology, Lawrence Berkeley National Lab, Berkeley, CA, an old Group image
- Youngho Seo, Department of Radiology and Biomedical Imaging, UCSF
- Marcus Hohlmann, Physics and Space Sc. Department, FIT
Acknowledgements
- US National Institute of Health
- US National Science Foundation
- US Department of Homeland Security
- Peter Belhumeur, Columbia U
Some university rankings
In the end I became fatigued with working on complete abstractions, and wanted to dip my hands into some real data, as I was doing during my early infatuation with computers. Mathematical or scientific computing started attracting me more. With a sabbatical at Lawrence Berkeley National Lab I was introduced to inverse problems over noisy nuclear imaging data. Medical image processing and management became my staple, but once again, on spatio-temporal data or dynamic images. Thanks to a number of extraordinary collaborators and students, our learning curve in this new area is steady.
Jorney continues. As the next stage of the above works, we are now involved with radiomics, asking - how to make prognosis from some 3D med-images of patients? Obviously, this is a machine learning task and our hands are dirty with tools like convolutional neural networks. The field is exploding and our heads whirl with recent new exotic works in the field. However, is there a role for topology in all these? Can studying topological invariances of images enhance learning capbility and efficiency? Are our works in spatio-temporal knowledge representation have some value in this direction?
So, here I am! But, stay tuned for more, in case you are tracking me :)
A disclaimer: I am not the Debasis Mitra, who works on distributed computing, now in Columbia University.
Projects
INVERSE PROBLEMS
SPATIO-TEMPORAL CONSTRAINT REASONING
Another question we have started asking recently (with a colleague) is whether a temporal reasoning approach may be helpful in timed-automata based model checking. The latter is a useful tool for verifying a designed system and a significant amount of literature exist on theoretical and practical problems.
CAPSTONE PROJECT: PREDICTING BRAIN SEIZURE WITH A MOBILE DEVICE
PAST PROJECT: COMPUTATIONAL MOLECULAR BIOLOGY
PAST PROJECT: KNOWLEDGE MANAGEMENT
PROTO-PROJECTS
In a similar line, can an NN be trained to detect a sequence of numbers, e.g., Fibonacci number? Will it be able to learn a generating function with pole or singularity? Can a generative NN do Fourier transform?
Publications
Nuclear Imaging
- ìReconstruction of 4-D Dynamic SPECT Images From Inconsistent Projections Using a Spline Initialized FADS Algorithm (SIFADS).î Mahmoud Abdalah, Rostyslav Boutchko, Debasis Mitra, Grant T. Gullberg. IEEE Transactions in Medical Imaging, 34(1): 216-228, 2015.
- ìClustering Initiated Factor Analysis (CIFA) Application for Tissue Classification in Dynamic Brain PET.î Rostyslav Boutchko, Debasis Mitra, Suzane Baker, William Jagust, and Grant T. Gullberg. Journal of Cerebral Blood Flow & Metabolism ñ Nature, doi:10.1038/jcbfm.2015.69, 2015.
- ìHigh Performance Fully 3D and 4D Image Reconstruction in SPECT Using a Big Data Analytic Tool Running on a Supercomputer.î Huang S-Y., Lee J.H., Pan H., Boutchko R., Shrestha U., Gullberg G.T., Mitra D., Yao Y., and Seo Y. The 13th International Meeting on Fully Three-Dimensional Image Reconstruction in Radiology and Nuclear Medicine (Fully-3D), Newport, Rhode Island, USA, 2015.
- ìParallelization of Iterative Reconstruction Algorithms in Multiple Modalities,î D. Mitra, H.Pan, Fares Alhassen, and Youngho Seo. Proc. IEEE Nuclear Science Symposium and Medical Imaging Conference, Seattle, WA, 2014.
- "SinoCor: Sinogram Level Motion Correction in SPECT", Debasis Mitra, Daniel Eiland, Rostyslav Boutchko, Grant T. Gullberg. Lawrence Berkeley National Laboratory Software Licensee CR-3016, 2013
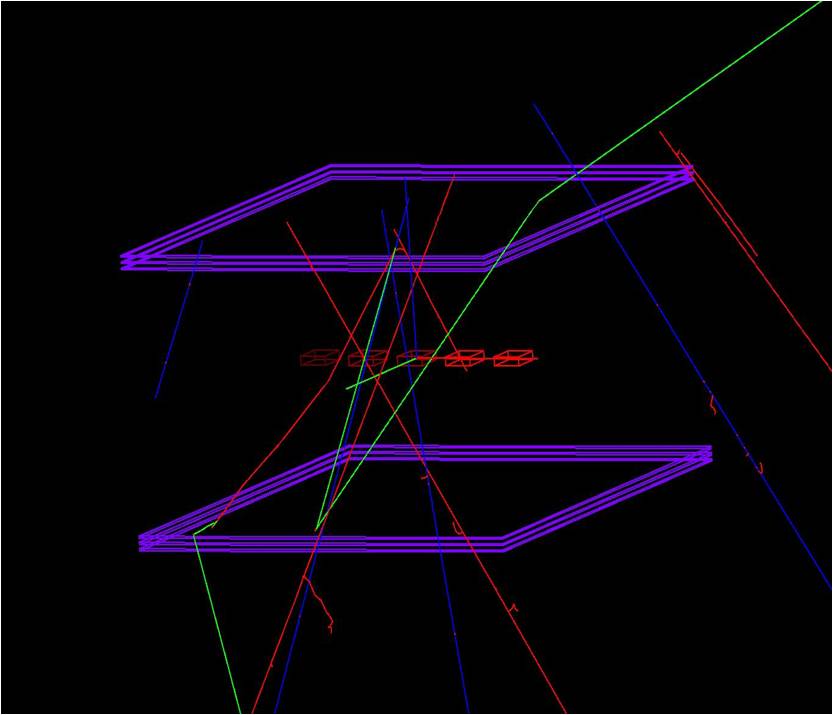
Muon Tomography
- "A Volume Clearing Algorithm for Muon Tomographyî D. Mitra, K. Day, and M. Hohlmann. Proceedings of IEEE Nuclear Science Symposium and Medical Imaging Conference, Seattle, WA, 2014.
- "Imaging of high-Z material for nuclear contraband detection with a minimal prototype of a muon tomography station based on GEM detectors." Nuclear Instruments and Methods in Physics Research A, , Kondo Gnanvo, Leonard V. Grosso III, Marcus Hohlman, Judson B. Locke, Amilkar Quintero, and Debasis Mitra. 652 (2011) 16ñ20
- “GEANT4 Simulation of a Cosmic Ray Muon Tomography System with Micro-Pattern Gas Detectors for the Detection of High-Z Materials,” "Transactions on Nuclear Science", Vol. 56, No. 3, June 2009, Marcus Hohlmann, Patrick Ford, Kondo Gnanvo, Jennifer Helsby, David Pena, Richard Hoch, and Debasis Mitra. [PDF]
- “Muon Tomography Algorithms for Nuclear Threat Detection,” "Lecture Notes in Artificial Intelligence" series, Springer Verlag, 2009, Richard Hoch, Debasis Mitra, Marcus Hohlman, and Kondo Gnanvo. [PDF]
- “Performance Expectations for a Tomography System Using Cosmic Ray Muons and Micro Pattern Gas Detectors for the Detection of Nuclear Contraband,” in Proc. of the IEEE Nucl. Sci. Symp. 2008, Dresden, Germany, pp. 1278-1284, IEEE Cat. CFP08NSS-CDR, ISBN 978-1-4244-2715-4, ISSN 1082-3654, e-Print: arXiv:0812.1007, Kondo Gnanvo, Patrick Ford, Jennifer Helsby, Richard Hoch, Debasis Mitra, and Marcus Hohlman. [PDF]
Spatio-temporal Reasoning
- “Explanation Generation over Temporal Interval Algebra,”
in Qualitative Spatio-Temporal Representation and Reasoning: Trends and Future Directions. Shyamanta M. Hazarika (Editor), Information Science Publishing, ISBN: 1616928689, August 1, 2010.
Debasis Mitra and Florent Launay. [PDF] - “Spatial-reasoning for Agents in Multiple Dimensions,”
Journal of Universal Computer Science, vol. 8, no. 8, pp. 774-791, August, 2002
Debasis Mitra, and Gerard Ligozat. [PDF] - “Spatial and Temporal Reasoning: Beyond Allen's Calculus,” Gerard Ligozat, Debasis Mitra and Jean-Francois Condotta, AI Communications (European journal on Artificial Intelligence), Vol. 17, no. 4, pp 223‚àö‚â•233, 2004.
- “Modeling and Reasoning with Star Calculus: An Extended Abstract,” Debasis Mitra, Proc. of Eighth International Symposium on AI and Math, January 2004, Ft. Lauderdale, Florida. http:/www.cs.fit.edu/~dmitra/Pub/MitraAiMath04CamRdy.pdf
- “Qualitative Direction Calculi with Arbitrary Granularity,” Jochen Renz and Debasis Mitra, Pacific Rim Conference on AI (PRICAI), 2004.
- "Characterization of Temporal Sequences in Geophysical Databases," Arie Shoshani, Preston Holland, Janet Jacobsen and Debasis Mitra, Proc. of the Statistical and Scientific Database Management (SSDBM) conference, Sweden, 1996.
Mathematical Physics
- “The Lorentz group in oscillator realization III - the group SO(3,1),” D. Basu and D. Mitra, Journal of Mathematical Physics, Vol 22, p 946, Am. Inst. Phys., 1981.
- “The Lorentz group in oscillator realization II - integral transform and matrix elements of SO(2,1),” D. Basu and D. Mitra, Journal of Mathematical Physics, Vol 21, p 636, Am. Inst. Phys., 1980.
Data & Knowledge Management
- “ReMI: An Object-relational Image Database for Nuclear Medicine Research,” Boutchko, Rostyslav; Fernandes, Antall; Pan, Hui; Abdalah, Mahmoud; Giannakidis, Archontis; Boswell, Martin; Mitra, Debasis; Gullberg, Grant T. Annual Conference of Society of Nuclear Medicine, Vancouver, BC, Canada, 2013.
- “A Data Management System with Web Interface for Pre-clinical Multi-modality Imaging: ReMI,” Mitra, Debasis ; Pan, Hui; Abdalah, Mahmoud; Boutchko, Rostyslav; Boswell, Martin; and Gullberg, Grant T. The Sixth World Molecular Imaging Congress, Savannah GA., 2013.
- “Three generations of research in computational creativity and beyond,” in Association for Advancement of Artificial Intelligence Spring Symposium on Creative Intelligent Systems (AAAI Tech Report). (Eds.) Ventura, D., Maher, M. L., and Colton, S.; Stanford, California, March 2008. Debasis Mitra.
- “Data cleaning and enriched representations for anomaly detection in system calls,” G. Tandon, P. Chan, and D. Mitra. Book Chapter in Machine Learning and Data Mining for Computer Security: Methods and Applications, M. Maloof (editor), Springer, 2005.
Computatinal Molecular Biology
- “Correlogram-based method for comparing biological sequences,” Debasis Mitra, Gandhali Samant and Kuntal Sengupta. Springer Verlag Lecture Notes on AI, 2006.

Life starts from today, always!
Florida Istitute of Technology,
Melbourne, Florida 32901
I am not avalable during Summer and Fall 2018
dmitra at cs.fit dot edu
Phone: 321-674-7737
Fax: 321-674-7046